Introduction
Knowledge Graphs (KGs) are advanced forms of networks that capture the semantics of the constituent entities and the interactions among them. They facilitate ontology-driven data consolidation via integration/harmonization of heterogeneous data and serve as a graphical database. Such KGs in place have the potential to answer complex queries and form the basis of domain-specific analyses. In context of biomedicine and life sciences, KGs represent disease-associated biological and pathophysiological phenomena by systematically assembling various inter-related entities such as proteins and their biological processes, molecular functions and pathways, chemicals and their mechanism of actions and adverse effects and so on. They have been deployed in several use cases and downstream analyses related to healthcare, pharmaceutical and clinical settings. However, the process of creating KGs is expensive and time-consuming because it requires a lot of manual curation. Moreover, machine-aided methods such as text-mining workflows and Large Language Models (LLMs) have their own shortcomings and are improving gradually.
This showcase introduces a fully automated workflow, namely Knowledge Graph Generator (KGG), for creating KGs that represent chemotype and phenotype of diseases. The KGG embeds underlying schema of curated public databases to retrieve relevant knowledge which is regarded as the gold standard for high quality data. The KGG is leveraged on our previous contributions to the BY-COVID project where we developed workflows for identification of bio-active analogs for fragments identified in COVID-NMR studies (Berg, H et al.) and representation of Mpox biology (Karki, R et al.). The programmatic scripts and methods for KGG are written in python (version 3.10) and are available on GitHub.
Who is the showcase intended for?
The KGG is developed for a broad spectrum of researchers and scientists, especially for those who are into pre-clinical drug discovery, understanding disease mechanisms/comorbidity and drug-repurposing. Although KGG is a programmatic tool, it comes with a user-friendly interface to take just a couple of input from a user to run the underlying scripts and methods. Therefore, it is designed to enable researchers with minimal knowledge of programming to generate KGs at ease. The computer scientists can make maximum advantage of the workflow by modifying the scripts according to their needs.
What is the showcase?
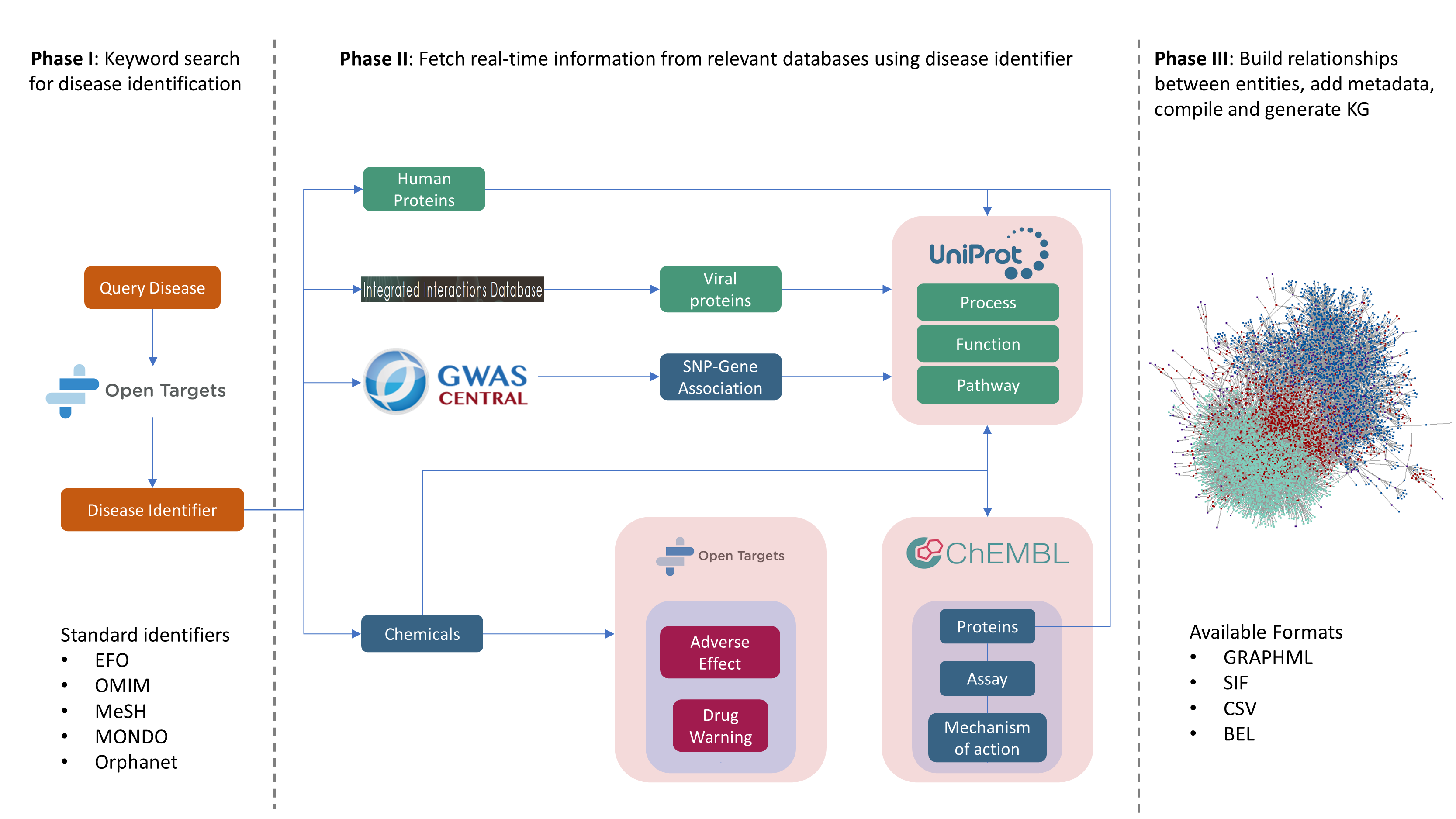
Figure 1. A schematic representation of the KGG workflow depicting its three phases. The python-based workflow fetches real-time knowledge from curated databases and uses the OpenBEL framework (Slater, T) to systematically encode the knowledge and relevant metadata.
The automated workflow creating disease-specific KGs is subdivided into three phases and are described below:
Phase I: Disease lookup and identification - The KGG workflow uses standard disease identifiers from widely accepted ontologies such as EFO, OMIM, MeSH, MONDO and so on. Therefore, the identification of a proper disease identifier for a specific disease is the foremost task in the workflow. In order to facilitate this task, we have designed KGG in such a way that the users can search disease names as keywords which are eventually passed as queries to the Open Target Platforms’s API. This step of the KGG workflow is termed as disease lookup which yields a list of diseases and identifiers closest to the keyword search. The users are then prompted to identify their disease of interest and the process of generating a KG can be initiated by using the corresponding identifier.
Phase II: Real-time knowledge retrieval - The identified disease identifier from Phase I is used as a query for curated databases to retrieve relevant disease associated knowledge in real time. This is achieved by embedding the APIs of OTP, ChEMBL, UniProt, Integrated Interaction Database (IID) and GWAS Central into our programmatic scripts and methods.
Phase III: KG compilation and generation - The retrieved knowledge from Phase II is stored as semantic triples (i.e., subject-predicate-object) using OpenBEL framework, which are both human and computer-readable. The language enables systematic representation of biological and molecular interactions by enforcing usage of standard ontologies. The implementation was performed using the open-source PyBEL framework. It is a resource developed to help with triples formation, meta-data annotation, data parsing, validation, compilation and visualization of KG. It also offers a wide-range of functions to explore, query, and analyze KGs. The KGs can be exported to various standard formats such as json, csv, sql, graphml, and Neo4j, allowing comparison and integration with other KGs.
What can you use the tool for?
The KGG serves the purpose of creating disease-specific KGs rapidly and swiftly. To our knowledge, this is the first workflow that enables it. One specific use case of KGG is to get the knowledge of important proteins and their functions within the context of any possible outbreak of infectious diseases. Moreover, researchers can also quickly identify drugs in different clinical trial phases of those diseases. Similarly, KGG can also help researchers to design their experiments as it incorporates knowledge of previously reported active assays. In addition to the main goal of creating automated workflows for KGs, we have also developed workflows to compare disease-specific KGs, shared entities between KGs from other resources, retrieve PDB structures of proteins and explore drug-likness of chemicals. Since KGs have many applications, they can also be of interest to bioinformaticians and chemoinformaticians for defining their own use case or deploying machine-learning methods.
Acknowledgments
Tool Development: Reagon Karki (Data Scientist, Fraunhofer ITMP/EU-OpenScreen), Andrea Zaliani (Data Scientist, Fraunhofer ITMP/EU-OpenScreen), Yojana Gadiya (Data Scientist, Fraunhofer ITMP)
Additional help: Philip Gribbon (Scientific project head/Director, Fraunhofer ITMP/EU-OpenScreen)
Support
This showcase was supported by Horizon Europe’s BY-COVID project [101046203] and EOSC Future Project [101017536].
More information
Skip tool tableTools and resources on this page
| Tool or resource | Description | Related pages | Registry |
|---|---|---|---|
| OpenBEL | The OpenBEL Framework is an open-platform technology for managing, publishing, and using biological knowledge represented using the Biological Expression Language (BEL). | ||
| PyBEL | Pure Python package for parsing and handling biological networks encoded in the Biological Expression Language (BEL). | Tool info |

